umxModify & umxCompare
Why make a new model when you can update one you already have?
This tutorial covers using umxModify.
In umx, models can be modified. Want to free a path? Go ahead - no need to start from scratch. You can add, or remove most things, and re-run, and compare in one call.
umxModify is a convenience function to add, set, or drop parameters, re-name, and re-run a model.
For what follows, we’ll need a model, so:
manifests = c("mpg", "disp", "gear")
m1 = umxRAM("mpg_model",data = mtcars, type = "cov",
umxPath(c("disp", "gear"), to = "mpg"),
umxPath("disp", with = "gear"),
umxPath(var = manifests)
)
Now show a summary.
umxSummary(m1, std = TRUE)
plot(m1, std= TRUE)
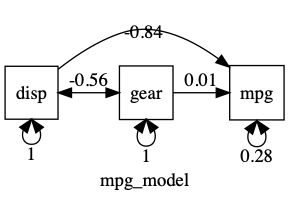
Modify a model
Fundamentally, modeling is in the service of understanding causality and we do that primarily via model comparison: Better theories fit the data better than do worse theories.
So, “do more gears give better miles per gallon”? In graph terms, this is asking, “is the path from to gear to mpg significant?” There are two ways to test this with umx.
1. Overwriting a path with OpenMx’s mxModel function
Using OpenMx’s mxModel function, new paths over-write existing paths. So adding a new path, with value fixed to zero will accomplish our goal:
m2 = mxModel(m1, umxPath("gear", to = "mpg", fixedAt = 0))
m2 = mxRun(m2)
You can then compare the two models
umxCompare(m1, m2)
2. Use umxModify() and labels
The easiest and most powerful way to modify models is with umx’s umxModify function.
For example, this one-liner drops the path “gear_to_mpg”, tests the effect, and returns the updated model:
m2 = umxModify(m1, update = "gear_to_mpg", name = "drop effect of gear", comparison = TRUE)
And viola:
| name | Estimate | SE |
|---|---|---|
| disp_to_mpg | -0.041 | 0.005 |
| mpg_with_mpg | 9.911 | 2.478 |
| disp_with_disp | 14880.775 | 3720.133 |
| disp_with_gear | -49.215 | 17.914 |
| gear_with_gear | 0.527 | 0.132 |
χ²(1) = -0.03, p = 1.000; CFI = 1.021; TLI = 1.063; RMSEA = 0
| Model | EP | Δ -2LL | Δ df | p | AIC | Δ AIC | Compare with Model |
|---|---|---|---|---|---|---|---|
| mpg_model | 6 | -0.047875 | 0 | ||||
| mpg_model | 5 | 0.0145592 | 1 | 0.904 | -2.033316 | -1.9854408 | mpg_model |
More advanced options in umxModify
- Pattern matching with regex So far we’ve seen update by matching a label:
fit2 = umxModify(fit1, update = "Cs", name = "newModelName")
A powerful additional feature of umxModify is regular expressions. These let you drop collections of paths by matching patterns.
So, to drop all paths with labels matching “gear_to_” followed by any characters, you could say:
m2 = umxModify(m1, regex = "gear_to_.*", name = "drop all paths from gear", comparison = TRUE)
nb: This is particularly powerful updating twin models with many paths to drop.
Regular expressions are very powerful.
fit2 = umxModify(fit1, update = "C[sr]", regex = TRUE, name = "drop_Cs_andCr")
You can also update a model by passing in new objects to be added to the model, like a CI request:
fit2 = umxModify(fit1, update = mxCI("Cs"), name = "withCI")
Also, we seen that you can request the model comparison summary at the same time:
To run umxCompare() and report on the old and new models, just set “comparison = TRUE”
Additional options for umxModify
free and value
By default, matched labels will be dropped, i.e., “free = FALSE value = 0”
To free (instead of fix) a variable, set free = TRUE.
To set the value of the matched paths to value X (instead of the default 0), set value = X.
To update only parameters that are already free, set freeToStart = TRUE. This defaults to NA - i.e, state is ignored.
more than one thing
Most parameters can take more than one input. To drop two paths, just offer up two labels in update.
m2 = umxModify(m1, update = c("path1", "path2"), name = "drop 2 paths")
tryHard
If you have a hard model to fit, you might want the optimizer to try extra hard. Just set tryHard = “yes”:
m2 = umxModify(m1, regex = "gear_to_.*", name = "drop paths from gear", tryHard = "yes")
update a label
If you want to update a label, you can do so as follows: In this case, setting to labels to a single newlabel
m2 = umxModify(m1, update = c("gear_to_mpg", "disp_to_mpg"), newlabels = "inputs_to_mpg", name = "equated_paths", tryHard = "yes")
note: this updates the model in raw units: standardized effects will likely differ. Best to work in standardized data for this sort of change.
Confidence Intervals
As in mxRun, you can set intervals = TRUE to run confidence intervals (see mxRun)
m2 = umxModify(m1, update = "gear_to_mpg", name = "no gear", intervals = TRUE)
note 1: For this to work, there have to be mxCIs in your model. See the post on mxConfint to learn more about confidence intervals.
note 2: SEs are often quicker and easier and are shown in the summary.