umxPath
“Staying on the (Wright) path”
umxPath is a core umx function. It facilitates path specification, streamlines scripts, and increases readability.
Let’s start using umxPath to specify a CFA model with a slight increase in complexity: two linked latent factors forced to be the same:
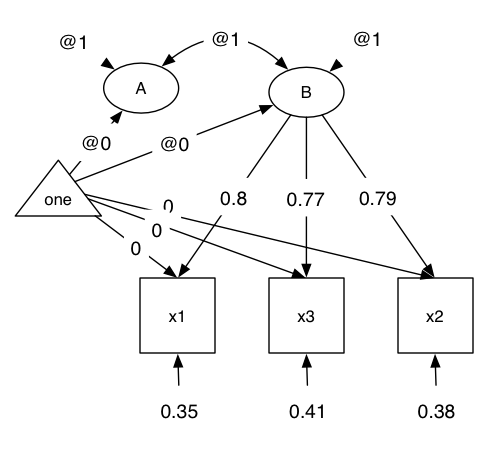
We’ll also use umxRAM to give us automagical parameter labeling and good start values.
# 2. set up some handy lists for the built-in myFADataRaw data set
manifests = paste0("x", 1:3) # "x1" "x2" "x3"
latents = c("A", "B")
m1 <- umxRAM("m1", data = myFADataRaw,
# 1-headed paths from latent B to each of the manifests
umxPath("B", to = manifests),
# A slightly silly unit-covariance between latent 1 & 2
umxPath("A", with = "B", fixedAt = 1),
# free mean and variance for the manifests and standardized latents.
umxPath(v.m. = manifests),
umxPath(v1m0 = latents)
)
plot(m1, showFixed = TRUE)
Shortcuts
You just learned two shortcuts offered by umxPath: The “v1m0” and “v.m.” options are short for “variance @1 and mean @ zero” and v.m. is show for “variance free, mean free” respectively.
The intuition for v.m. is v for variance, m for means, and dot for “any value”
We could break this out into 4 lines if desired:
umxPath(means = manifests),
umxPath(var = manifests),
umxPath(means = latents, fixedAt = 0),
umxPath(var = latents, fixedAt = 1),
The umxPath terms for connecting variables are shown below, along with their mxPath equivalents:
| New Term | Example | mxPath Equivalent |
|---|---|---|
| var | var = “X” | from = “X”, arrows = 2 |
| with | “X”, with = “Y” | from = “X”, to = “Y”, arrows = 2 |
| cov | cov = c(“X”,”Y”) | from = “X”, to = “Y”, arrows = 2 |
| means | means = c(“X”,”Y”) | from = “one”, to = c(“X”,”Y”) |
| v1m0 | v1m0 = “X” | requires 2 lines setting mean & var |
| v0m0 | v0m0 = “Y” | requires 2 lines setting mean & var |
| v.m. | v.m. = “Y” | requires 2 lines setting mean & var |
| defn | defn = “A”, label=”data.Y” | create a latent var@0 mean fixed, with data.var as label |
| unique.bivariate | unique.bivariate = c(“X”,”Y”, “Z”) | from = c(“X”,”Y”, “Z”), connection = “unique.bivariate”, arrows = 2 |
| unique.pairs. | unique.pairs = c(“X”,”Y”, “Z”) | from = c(“X”,”Y”, “Z”), connection = “unique.pairs”, arrows = 2 |
| Cholesky | Cholesky = c(“X”,”Y”), to = c(“a”, “b”) | a complex series of paths |
unique.bivariate
unique.bivariate is a time saver and error-preventer. It allows a 1-line specification of bivariate paths between all distinct pairs of variables.
So to create paths “A↔B”, “B↔C”, and “A↔C”, you can say just:
umxPath(unique.bivariate = c('A', 'B', 'C')
and get back the equivalent of
umxPath("A", with = "B")
umxPath("A", with = "C")
umxPath("B", with = "C")
Cholesky
Cholesky has a page of its own as this feature moves us into twin modeling.
New words for setting values
The new words for setting values are shown below, along with their mxPath equivalents
| New term | mxPath Equivalent |
|---|---|
| fixedAt = 1 | free = FALSE, values = 1 |
| firstAt = 1 | free = c(FALSE, TRUE, TRUE), values = c(1, NA, NA) |
| freeAt = .6 | free = TRUE, values = .6 |
firstAt saves you laboriously counting out and keeping check of linked lists of free and value items. freeAt allows you to manually set a starting value.
Connection types
umxPath still lets you do anything mxPath does, and one cool thing about mxPath is it has many connection types.
I’ve exposed just unique.bivariate as a umxPath parameter, but the complete list of options is
- “single”
- “all.pairs”
- “all.bivariate”
- “unique.pairs”
- “unique.bivariate”
Default value is “single”.
Values and labels
You might have noticed that we can say things like
umxPath(c("A", "B", "C"), values = 1)
Two things are happening here. First, the values field is being “recycled” for each of the three paths being created here, so the three paths will be started at 1. Second is the from field is being recycled into the to field. so:
umxPath(from = "A")
is the same as:
umxPath(from = "A", to = "A")
QuickLook: look often, run once
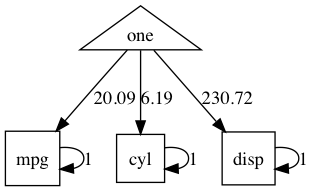
Tip: plot() is a great way to see what you are doing in a model as you build it: look often build once.
plot(umxRAM("tim", umxPath(c("mpg", "cyl", "disp"), value=1), data=mtcars, run=F))
# hmmm single-headed arrows... should use "var = "